Lantrell Stewart, “Detection of Tissue Factor Expression on Transfected Cells”
Mentor: Julie Oliver, Biological Sciences
We detected human tissue factor (TF) expression by direct and indirect methods on transfected clones of the 300.19 mouse pre-B cell line. TF is a trans-membrane protein receptor responsible for initiating coagulation. It is constitutively expressed on cells outside the vasculature, and under pathological conditions, can be expressed on endothelial cells and some leukocytes. The cell lines used in this experiment contained fusion proteins of TF with an attached fluorophore, either singly transfected TF-mTurquoise (mTq), singly transfected TF-super yellow fluorescent protein 2 (SYFP2), or lines co-transfected with both. We employed flow cytometry analysis of the attached fluorophore for direct TF detection. Using a FACSCalibur flow cytometer, we were able to quantify expression in lines transfected with TF-SYFP2. However, the fluorescence capabilities of the instrument do not allow direct analysis of TF-mTq expression. Therefore, we used indirect detection to quantify TF expression in the mTq- and co-transfected lines. Cells were labeled with a mouse anti-human TF monoclonal antibody and a goat anti-mouse IgG antibody conjugated with Alexa Fluor 647. We hypothesized that in co-transfectants, about half the total TF detected by the indirect method could be explained by the direct SYFP2 flow cytometry signal. This was our actual observation in every co-transfectant tested. We were able to confirm the flow cytometry results by qualitative fluorescence microscopy imaging using filter sets appropriate for all three fluorophores used (mTq, SYFP2, and Alexa Fluor 647). Therefore, using a combination of detection methods has provided us great flexibility in analysis of transfected TF expression.
Click the thumbnail below to open the full sized poster in a new tab.
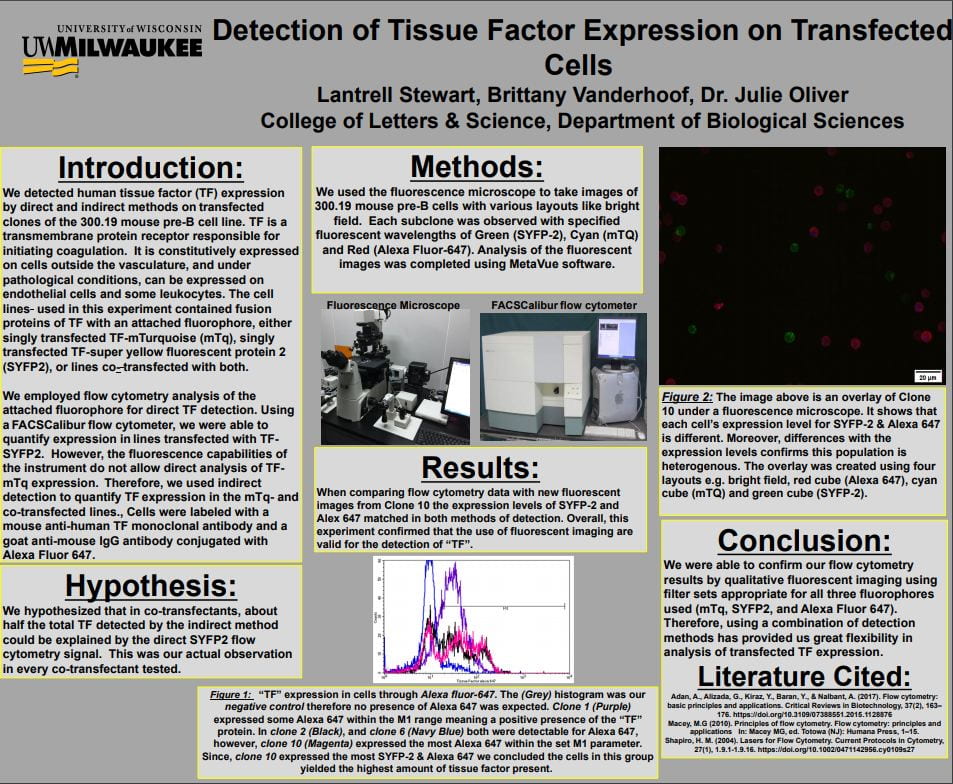
The layout of the different sections was nice, but the project was hard to follow for anyone outside the area of research (the introduction contained too much specialized vocabulary, and would have benefitted from making it more accessible to a general audience). The introduction also contained a lot of information that would have been better placed in methods.